|
| | form ($form, &$form_state) |
| |
| | formValidate ($form, &$form_state) |
| |
| | run () |
| |
| | formSubmit ($form, &$form_state) |
| |
| | __construct (array $configuration, $plugin_id, $plugin_definition, ChadoConnection $connection, ?Messenger $messenger=NULL, ?TripalLogger $logger=NULL, ?TripalFileRetriever $fileretriever=NULL, ?TripalBackendPublishManager $publish_manager=NULL,) |
| |
| | getChadoConnection () |
| |
| | startTransactions () |
| |
| | addAnalysis ($form, &$form_state) |
| |
| | postRun () |
| |
| | __construct (array $configuration, $plugin_id, $plugin_definition, ?Messenger $messenger=NULL, ?TripalLogger $logger=NULL, ?TripalFileRetriever $fileretriever=NULL, ?TripalBackendPublishManager $publish_manager=NULL,) |
| |
| | describeUploadFileFormat () |
| |
| | setJob ($job) |
| |
| | rollbackTransaction (string $stage) |
| |
| | createImportJob ($run_args, $file_details=[]) |
| |
| | load ($import_id) |
| |
| | submitJob () |
| |
| | prepareFiles () |
| |
| | cleanFile () |
| |
| | setArguments (array $arguments) |
| |
| | getArguments () |
| |
|
| | retrieveLineage ($root_taxon=NULL) |
| |
| | initTree ($tree_name) |
| |
| | removeTree ($tree_name) |
| |
| | rebuildTree ($root_taxon=NULL) |
| |
| | rebuildTreeOrganismLineage ($organism, $root_taxon, &$tree, &$lineage_good) |
| |
| | queryPhylonode (int $phylotree_id, string $node_name, bool &$lineage_good, array &$lineage_nodes) |
| |
| | getPhylonode (int $phylotree_id, int $organism_id) |
| |
| | addOrganismNode ($organism, $root_taxon=NULL) |
| |
|
| addTaxonomyNode (&$tree, $node, $lineage_elements) |
| |
| | trimLineage ($lineage, $root_taxon=NULL) |
| |
| | addBundleTypeProperty (string $pkey, int $record_id, string $property_table, string $termIdSpace, string $termAccession, string $value='', ?int $rank=NULL) |
| |
| | setTotalItems ($total_items) |
| |
| | addItemsHandled ($num_handled) |
| |
| | setItemsHandled ($total_handled) |
| |
| | setInterval ($interval) |
| |
| | xmlIsValid (string $xml) |
| |
|
| static | create (ContainerInterface $container, array $configuration, $plugin_id, $plugin_definition,) |
| |
◆ addOrganismNode()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::addOrganismNode |
( |
|
$organism, |
|
|
|
$root_taxon = NULL |
|
) |
| |
|
protected |
Adds an organism to the taxonomy tree using its lineage.
- Parameters
-
| $organism | An organism object. |
| $root_taxon | An optional taxon name for the root node if generating a sub-tree. |
- Returns
- int Status. 1 = added 2 = not added, because not part of $root_taxon (if specified) 3 = no lineage available for this organism
◆ form()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::form |
( |
|
$form, |
|
|
& |
$form_state |
|
) |
| |
◆ formSubmit()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::formSubmit |
( |
|
$form, |
|
|
& |
$form_state |
|
) |
| |
{Handles submission of the form elements.The form elements provided in the implementation of the form() function can be used for special submit if needed.
- Parameters
-
| array | $form | The form array definition. |
| \Drupal\Core\Form\FormStateInterface | $form_state | The form state object. |
}
Implements Drupal\tripal\TripalImporter\Interfaces\TripalImporterInterface.
◆ formValidate()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::formValidate |
( |
|
$form, |
|
|
& |
$form_state |
|
) |
| |
◆ getPhylonode()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::getPhylonode |
( |
int |
$phylotree_id, |
|
|
int |
$organism_id |
|
) |
| |
|
protected |
Retrieves a phylonode in a phylotree for an organism
- Parameters
-
| int | $phylotree_id | The phylotree to query |
| int | $organism_id | The organism to query |
- Returns
- object A phylonode object, or NULL if no match.
◆ initTree()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::initTree |
( |
|
$tree_name | ) |
|
|
protected |
Create the taxonomic tree in Chado.
If the tree already exists it will not be recreated.
- Exceptions
-
- Returns
- Returns the phylotree object.
◆ queryPhylonode()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::queryPhylonode |
( |
int |
$phylotree_id, |
|
|
string |
$node_name, |
|
|
bool & |
$lineage_good, |
|
|
array & |
$lineage_nodes |
|
) |
| |
|
protected |
Called by rebuildTreeOrganismLineage(), selects a phylonode and its properties.
We need to find the node in the phylotree for this level of the lineage, but there are a lot of repeats and we don't want to keep doing the same queries over and over, so we store the nodes we've already seen in the $lineage_nodes array for fast lookup.
◆ rebuildTree()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::rebuildTree |
( |
|
$root_taxon = NULL | ) |
|
|
protected |
Iterates through all existing organisms and builds the taxonomy tree.
The phloytree API doesn't support adding nodes to existing trees, only importing whole trees. So, we must rebuild the tree using the current organisms.
- Parameters
-
| string | $root_taxon | If specified, this taxon is the root of the tree, and any organisms not part of this taxon are excluded from the tree. |
◆ rebuildTreeOrganismLineage()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::rebuildTreeOrganismLineage |
( |
|
$organism, |
|
|
|
$root_taxon, |
|
|
& |
$tree, |
|
|
& |
$lineage_good |
|
) |
| |
|
protected |
Called by rebuildTree(), adds lineage nodes for one organism.
◆ removeTree()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::removeTree |
( |
|
$tree_name | ) |
|
|
protected |
Removes the chado.phylotree record. Used when tree generation is cancelled due to lack of any valid organisms.
◆ retrieveLineage()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::retrieveLineage |
( |
|
$root_taxon = NULL | ) |
|
|
protected |
Parses NCBI Taxonomy lineage details for organisms at this site. This is information previously generated in Chado by the taxonomy importer.
◆ run()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::run |
( |
| ) |
|
◆ trimLineage()
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::trimLineage |
( |
|
$lineage, |
|
|
|
$root_taxon = NULL |
|
) |
| |
|
protected |
Removes any part of a lineage above an optional root taxon, and returns the resulting lineage as an array.
- Parameters
-
| string | $lineage | The semicolon-delimited taxonomic lineage. If is lineageex it has colon-delimited parts. |
| string | $root_taxon | The root taxon, e.g. a family, or NULL. |
- Returns
- array $lineage_elements Lineage string exploded into an array, starting at root taxon if specified.
◆ $all_orgs
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::$all_orgs = [] |
|
protected |
Holds the list of all organisms currently in Chado.
◆ $phylotree
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::$phylotree = NULL |
|
protected |
The record from the Chado phylotree table that refers to this Taxonomic tree.
◆ $rank_cvterm_id
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::$rank_cvterm_id = NULL |
|
protected |
CV term id for local:rank
◆ $tree
| Drupal\tripal_chado\Plugin\TripalImporter\TreeGenerator::$tree = NULL |
|
protected |
The temporary tree array used by the Tripal Phylotree API for importing a new tree.
The documentation for this class was generated from the following file:
- tripal_chado/src/Plugin/TripalImporter/TreeGenerator.php
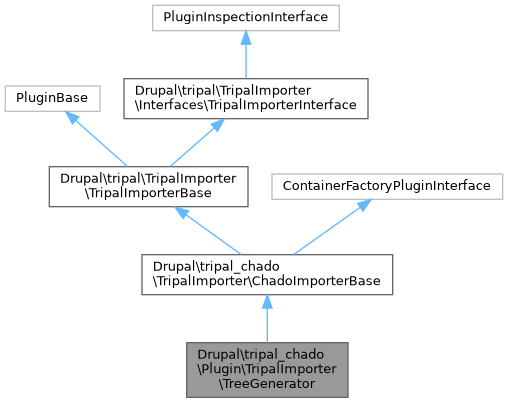
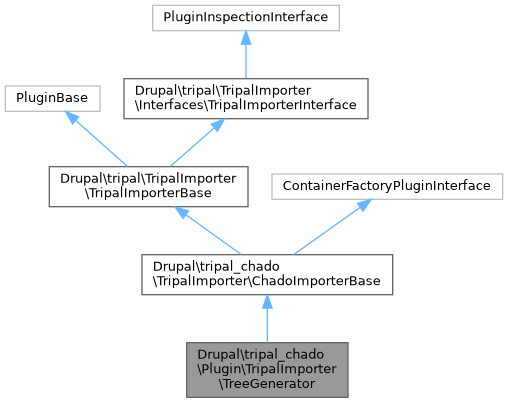
 Public Member Functions inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase
Public Member Functions inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase Public Member Functions inherited from Drupal\tripal\TripalImporter\TripalImporterBase
Public Member Functions inherited from Drupal\tripal\TripalImporter\TripalImporterBase Protected Member Functions inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase
Protected Member Functions inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase Protected Member Functions inherited from Drupal\tripal\TripalImporter\TripalImporterBase
Protected Member Functions inherited from Drupal\tripal\TripalImporter\TripalImporterBase Protected Attributes inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase
Protected Attributes inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase Protected Attributes inherited from Drupal\tripal\TripalImporter\TripalImporterBase
Protected Attributes inherited from Drupal\tripal\TripalImporter\TripalImporterBase Static Public Member Functions inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase
Static Public Member Functions inherited from Drupal\tripal_chado\TripalImporter\ChadoImporterBase